Figures & data
Figure 1. Wild-growing Tulipa species documented in Greece.
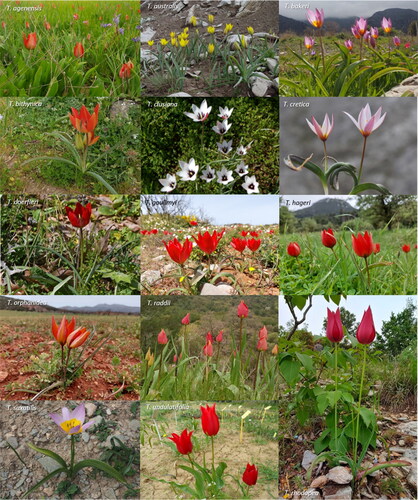

Figure 2. Phylogenetic tree of 34 Tulipa taxa with DNA sequences generated from wild-growing Greek tulip species (in green letters) and 22 retrieved from GenBank (in black letters). Sequence analysis was based on the ITS + psbA-trnH + trnL-trnF molecular markers and phylogenetic differences were estimated by using the Neighbor-Joining method. The evolutionary distances were computed using the Maximum Composite Likelihood method and are in the units of the number of base substitutions per site. Coding of samples and nomenclature of taxa as submitted in the GenBank database.
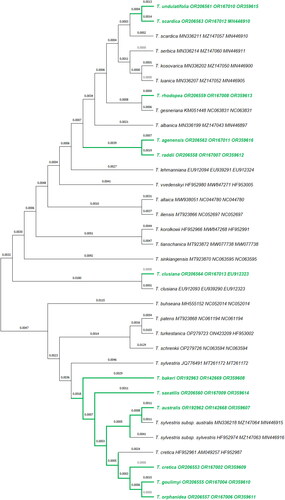
Table 1. Overview of nucleotide variations based on DNA sequence analysis of the psbA-trnH and trnL-trnF cpDNA molecular markers after comparison of 25 DNA sequences generated for the 15 wild-growing greek tulip species with 80 DNA sequences retrieved from GenBank (nomenclature of taxa as submitted in GenBank).
Table 2. Overview of nucleotide variations in specific loci based on DNA sequence analysis of the psbA-trnH and trnL-trnF cpDNA molecular markers after comparison of the studied specimens of the 15 wild-growing greek tulip species with DNA sequences retrieved from GenBank (YES/NO indicates the presence/absence of the nucleotide variation, and dash the absence of data for comparison).
Supplemental Material
Download Zip (1.4 MB)Data availability statement
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
