Figures & data
Figure 1. High-fat diet increased susceptibility to oral and systemic Listeria monocytogenes infection, but this increase was absent in mice treated with A. muciniphila.
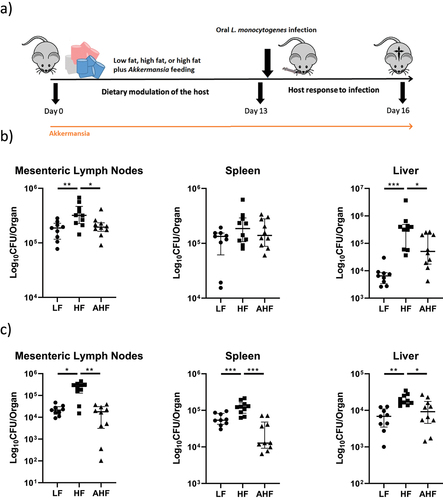
Figure 2. A. muciniphila reduces inflammatory expression and goblet cell numbers in the ileum while exacerbating hepatic immune expression.
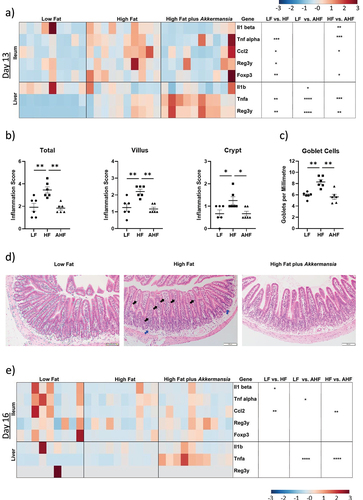
Figure 3. The protective effect of A. muciniphila does not appear to be driven by changes in the gut microbiota.
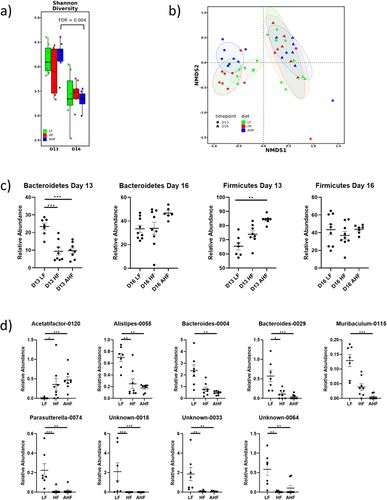
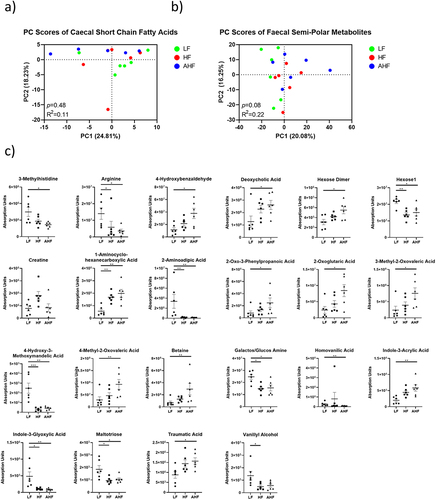
Figure 4. The protective effect of A. muciniphila does not appear to be driven by fecal or cecal metabolomes at day 13.
Data availability statement
All data are available on Zenodo at the following link https://doi.org/10.5281/zenodo.7590374.
