Figures & data
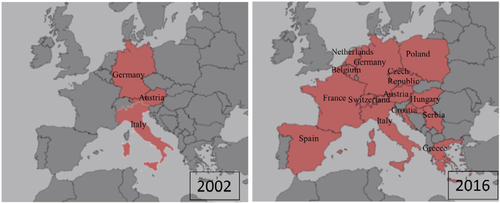
Figure 1 Usutu virus phylogram trees.
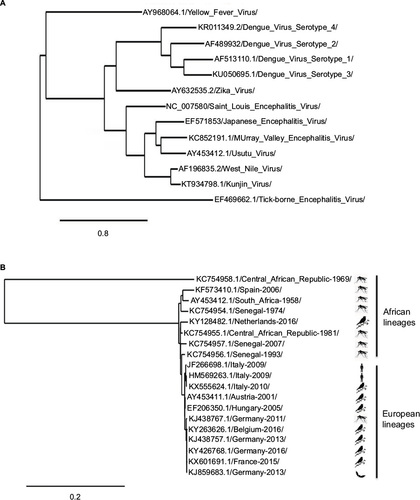
Figure 2 Schematic view of the genomic organization of USUV.
Abbreviations: C, capsid; E, envelope; ORF, open reading frame; prM/M, pre-membrane/membrane; USUV, Usutu virus; UTRs, untranslated regions.
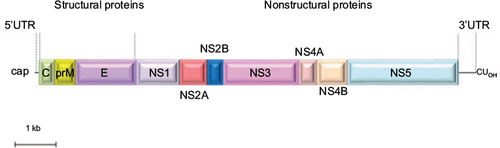
Figure 3 Transmission electron microscopy of USUV RSPs and virions.
Abbreviations: RSPs, recombinant subviral particles; TEM, transmission electron microscopy; USUV, Usutu virus; VP, vesicle packets; Vi, virions.
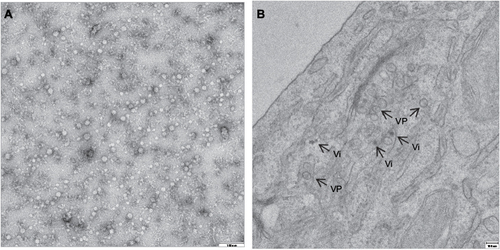
Figure 4 Confocal image of Vero cell transfected with the autophagosome marker GFP-LC3 plasmid and infected with USUV (red).
Abbreviation: USUV, Usutu virus.
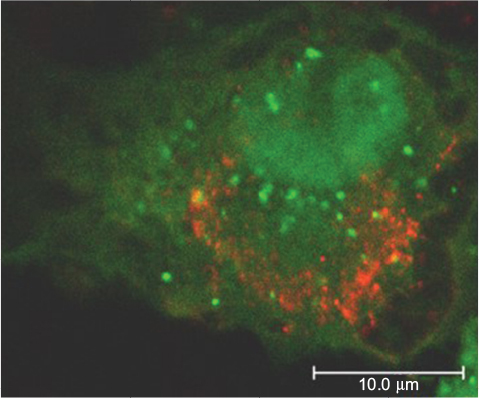
Table 1 Circulation of USUV among avian species in Europe